Building the AI Clinical Trial Matching Platform That Reduced Patient-to-Trial Time for a Global Pharma Leader
Manual patient screening across disconnected EHR and genomic systems made trial enrollment slow and inaccurate. Ideas2IT built an AI matching platform combining genomic, clinical, and histological data that reduced matching time from weeks to hours and gave clinical teams real-time trial visibility.

Client
Global pharma leader

Industry
Pharma & Life Sciences

Service
Artificial Intelligence
Data Engineering

Compliance
HIPAA · GDPR

Collaboration
MolecularMatch
01 Challenge
Clinical teams operated across disconnected EMRs, genomic labs, hospital chains, and insurance systems with no unified patient view. Matching ran on static rule sets that couldn't process NGS genomic data, ECOG scores, or resistance biomarkers. Every enrollment window missed was a delayed trial and a patient who didn't get in.
02 Solution
Ideas2IT built a HIPAA, GDPR, and 21 CFR Part 11-compliant AWS data lake with FHIR HL7-R4 as the canonical format across all source systems. On that foundation, working with MolecularMatch, the team built a precision matching engine combining genomic, clinical, and histological data, and an NLP query interface that gave clinical teams insight without BI tool dependency.
03 Outcome
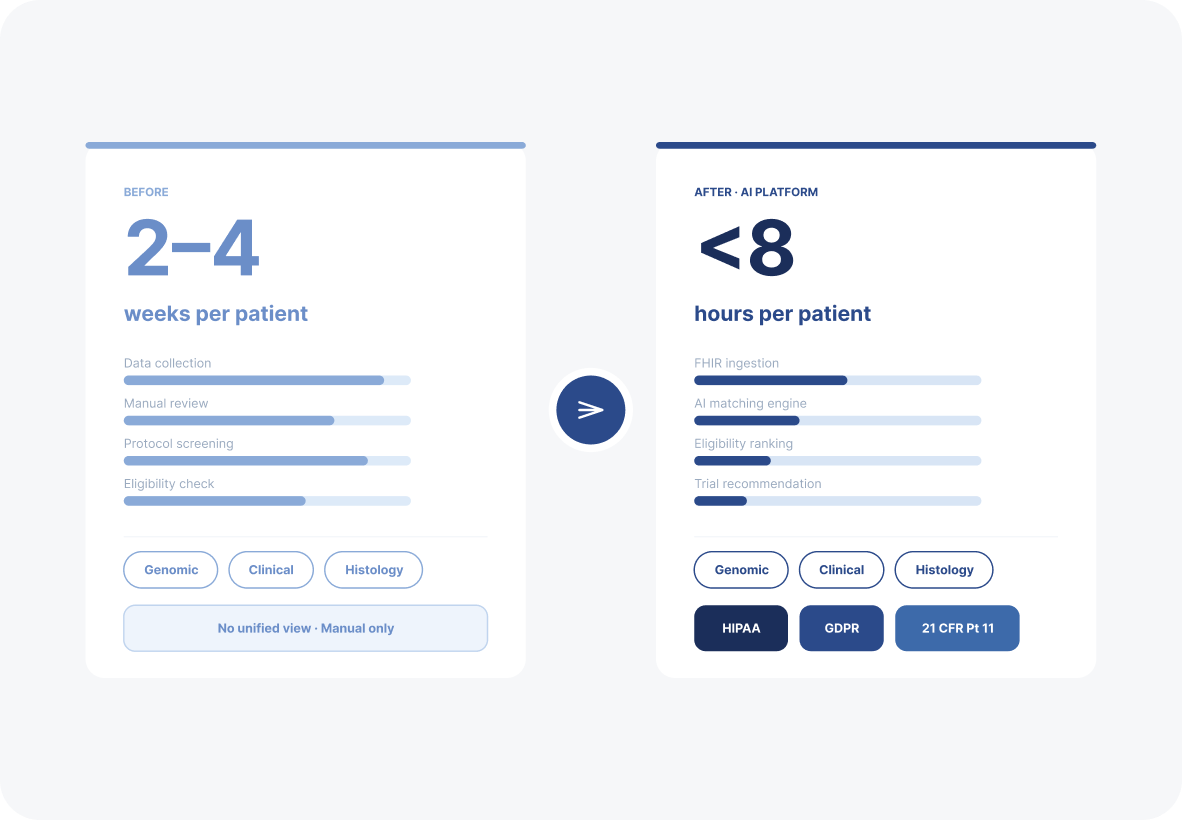
Patient-to-trial matching dropped from 2 to 4 weeks to under 8 hours. Protocol search time fell 60%. Decision-making speed increased 2.5x across clinical and business teams. The platform now powers real-time enrollment across oncology and immunotherapy trials.
Phase 01
Building the unified clinical data layer before matching could run
Compliant data foundation: FHIR ingestion, multi-region AWS data lake, and audit controls at every layer
The first constraint was format. Every source system hospital chains, EMRs, insurance providers, diagnostic labs produced clinical data in a different structure. FHIR HL7-R4 had to become the canonical format across all of them before any matching logic could operate.
Ideas2IT built
- A multi-region AWS data lake with an ingestion pipeline that converted Medication Statements, Diagnostic History, and clinical records from each source system into standardised FHIR.
- Role-based access controls, AES-256 encryption, and a full audit trail service embedded HIPAA, GDPR, and 21 CFR Part 11 compliance at every layer.
- Java microservices on SQS/SNS handled real-time data routing to subscribed applications, and dynamic task services on AWS ECS/EKS managed bulk FHIR operations at scale.
DELIVERABLES
- Multi-region HIPAA/GDPR-compliant AWS data lake
- S3-backed, jurisdictionally segregated FHIR HL7-R4 ingestion and transformation pipeline
- Real-time data routing layer
- Java microservices on SQS/SNS Bulk FHIR operation services
- Dynamic task services on AWS ECS/EKS Audit trail and compliance services
- HIPAA, GDPR, 21 CFR Part 11 Role-based access and AES-256 encryption framework
Phase 02
Reducing matching cycles from weeks to hours
AI matching engine and NLP interface: genomic precision and sub-8-hour enrollment windows
The matching engine required all three data layers together. Genomic alone produced false positives. Clinical context alone missed biomarker-driven exclusions.
Ideas2IT collaborated with MolecularMatch to build a proprietary engine that combined genomic, clinical, and histological data to rank trial eligibility in real time against ECOG performance scores, comorbidities, and resistance biomarkers. The trial database refreshed continuously, with REST APIs delivering ranked recommendations directly into care team systems.
On top of that, an NLP query interface translated plain English clinical questions into SQL, surfaced auto-generated visualisations, and gave non-technical users access to the full data estate without any BI tool. Matching time dropped from 2 to 4 weeks to under 8 hours.
DELIVERABLES
- Proprietary AI trial matching engine
- Genomic, clinical, and histological data combined MolecularMatch integration
- Clinical informatics matching layer
- Continuously refreshed trial database
- Real-time eligibility ranking REST API integration layer
- Trial recommendations into care team systems NLP query interface
- Plain English to SQL with auto-visualisation ECOG, biomarker, and comorbidity scoring
- Real-time eligibility filter layer
Phase 03
Bringing trial and treatment intelligence to clinicians
Clinical decision support platform: NCCN protocols, pathway authoring, and longitudinal patient view
The final layer had to close the gap between data and clinical decision. NCCN oncology protocols and institution-specific treatment pathways lived in separate systems, forcing clinicians to context-switch during protocol decisions.
Ideas2IT built an interactive flowchart interface that unified NCCN guidelines with custom institutional pathways, with smart search, source-linked transparency, and pathway authoring tools that clinical teams could configure without engineering support.
An NLP extraction layer pulled structured insights from unstructured medical records across each patient's timeline. Configurable biomarker tracking and integrated document access consolidated the clinical view into a single interface deployed across web and mobile. Protocol search time fell 60% and clinical decision speed increased 2.5x.
DELIVERABLES
- NCCN and institutional protocol unification
- Single interactive flowchart interface Smart protocol search with source-linked transparency
- Across oncology guidelines Custom pathway authoring tool
- Institution-configurable, no engineering required
- Longitudinal patient journey visualisation
- NLP extraction from unstructured medical records
- Structured insight layer Mobile and web deployment
- Single clinical interface across platforms
The Outcome
From 2-week manual screening to under 8 hours:
The matching time reduction from weeks to hours required a new data foundation, a new matching model, and a new interface layer built in sequence, each unlocking the next. The FHIR-standardised data lake made the matching engine possible. The matching engine made real-time eligibility ranking possible. The NLP interface made that precision accessible to clinical teams without technical barriers. That sequence is what produced the result.
